-Search query
-Search result
Showing 1 - 50 of 1,391 items for (author: sil & p)

EMDB-43246: 
TehA from Haemophilus influenzae purified in DDM
Method: single particle / : Catalano C, Senko S, Tran NL, Lucier KW, Farwell AC, Silva MS, Dip PV, Poweleit N, Scapin G

EMDB-43247: 
TehA from Haemophilus influenzae purified in GDN
Method: single particle / : Catalano C, Senko S, Tran NL, Lucier KW, Farwell AC, Silva MS, Dip PV, Poweleit N, Scapin G

EMDB-43248: 
TehA from Haemophilus influenzae purified in LMNG
Method: single particle / : Catalano C, Senko S, Tran NL, Lucier KW, Farwell AC, Silva MS, Dip PV, Poweleit N, Scapin G

EMDB-43249: 
TehA from Haemophilus influenzae purified in OG
Method: single particle / : Catalano C, Senko S, Tran NL, Lucier KW, Farwell AC, Silva MS, Dip PV, Poweleit N, Scapin G

PDB-8vi2: 
TehA from Haemophilus influenzae purified in DDM
Method: single particle / : Catalano C, Senko S, Tran NL, Lucier KW, Farwell AC, Silva MS, Dip PV, Poweleit N, Scapin G

PDB-8vi3: 
TehA from Haemophilus influenzae purified in GDN
Method: single particle / : Catalano C, Senko S, Tran NL, Lucier KW, Farwell AC, Silva MS, Dip PV, Poweleit N, Scapin G

PDB-8vi4: 
TehA from Haemophilus influenzae purified in LMNG
Method: single particle / : Catalano C, Senko S, Tran NL, Lucier KW, Farwell AC, Silva MS, Dip PV, Poweleit N, Scapin G

PDB-8vi5: 
TehA from Haemophilus influenzae purified in OG
Method: single particle / : Catalano C, Senko S, Tran NL, Lucier KW, Farwell AC, Silva MS, Dip PV, Poweleit N, Scapin G

EMDB-18664: 
Structure of the native microtubule lattice nucleated from the yeast spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

EMDB-18665: 
Structure of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

EMDB-18666: 
Structure of the y-Tubulin Small Complex (yTuSC) as part of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

PDB-8qv0: 
Structure of the native microtubule lattice nucleated from the yeast spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

PDB-8qv2: 
Structure of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D

PDB-8qv3: 
Structure of the y-Tubulin Small Complex (yTuSC) as part of the native y-Tubulin Ring Complex (yTuRC) capping microtubule minus ends at the spindle pole body
Method: subtomogram averaging / : Dendooven T, Yatskevich S, Burt A, Bellini D, Kilmartin J, Barford D
EMDB-41434: 
Bottom cylinder of high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L, Kirst H, Kerfeld CA
EMDB-41435: 
Central rod disk in C1 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L, Kirst H, Kerfeld CA
EMDB-41436: 
Central rod disk in D3 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L, Kirst H, Kerfeld CA
EMDB-41463: 
Synechocystis PCC 6803 Phycobilisome quenched by OCP, high resolution
Method: single particle / : Sauer PV, Sutter M, Cupellini L

EMDB-41475: 
Top cylinder bound to OCP from high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L
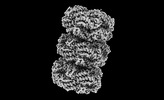
EMDB-41585: 
Rod from high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L

PDB-8to2: 
Bottom cylinder of high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L

PDB-8to5: 
Central rod disk in C1 symmetry of high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L

PDB-8tpj: 
Top cylinder bound to OCP from high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L

PDB-8tro: 
Rod from high-resolution phycobilisome quenched by OCP (local refinement)
Method: single particle / : Sauer PV, Sutter M, Cupellini L

EMDB-16901: 
CryoEM structure of 20S Trichomonas vaginalis proteasome in complex with proteasome inhibitor Salinosporamid A
Method: single particle / : Silhan J, Fajtova P, Boura E

PDB-8oix: 
CryoEM structure of 20S Trichomonas vaginalis proteasome in complex with proteasome inhibitor Salinosporamid A
Method: single particle / : Silhan J, Fajtova P, Boura E

EMDB-43658: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43659: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-43660: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vye: 
SARS-CoV-2 S (C.37 Lambda variant) plus S309, S2L20, and S2X303 Fabs
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vyf: 
SARS-CoV-2 S NTD (C.37 Lambda variant) plus S2L20 and S2X303 Fabs, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

PDB-8vyg: 
SARS-CoV-2 S RBD (C.37 Lambda variant) plus S309 Fab, local refinement
Method: single particle / : McCallum M, Veesler D, Seattle Structural Genomics Center for Infectious Disease (SSGCID)

EMDB-40208: 
Backbone model of de novo-designed chlorophyll-binding nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

EMDB-40209: 
Chlorophyll-binding region of de novo-designed nanocage O32-15
Method: single particle / : Redler RL, Ennist NM, Wang S, Baker D, Ekiert DC, Bhabha G

EMDB-17509: 
Cryo-EM structure of CAK in complex with inhibitor BS-181
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17510: 
Cryo-EM structure of CAK in complex with inhibitor BS-194
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17512: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0510-R
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17513: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0510-S
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17514: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0574
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17515: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0768
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17516: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0829
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17517: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0880 (ring-up conformation)
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17518: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0880 (ring-down conformation)
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17519: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0914
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17520: 
Cryo-EM structure of CAK in complex with inhibitor ICEC0943
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17521: 
Cryo-EM structure of CAK in complex with inhibitor dinaciclib
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17522: 
Cryo-EM structure of CAK with averaged inhibitor density
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17523: 
Cryo-EM structure of apo CAK
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17524: 
Cryo-EM map of inhibitor-bound CAK (Krios G4 performance comparison dataset)
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ

EMDB-17525: 
Cryo-EM map of inhibitor-bound CAK (Glacios G2 performance comparison dataset)
Method: single particle / : Cushing VI, Koh AF, Feng J, Jurgaityte K, Bahl AK, Ali S, Kotecha A, Greber BJ
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
